Rapid Cell-Free Production of Active Proteins
Development of an AI ready protein expression workflow to enable rapid activity screening of toxic proteins

Project overview
The challenge
- Proteins toxic to host cells are a challenge to produce.
- Multi-week timescales for protein production can delay downstream assays and are unsuitable for rapid iterative screening of proteins designed using AI methods.
Our solution
- Use of a cell free in vitro transcription and translation (IVTT) platform allows production of toxic proteins.
- Rapid single step purification of microgram yields.
- Protein activity confirmed by small molecule binding using Surface Plasmon Resonance (SPR).
Our impact
- Speed - from DNA to protein to SPR results within 48 hours.
- Purity and yield - suitable for initial SPR analysis .
- Medium throughput platform - suitable for screening of AI- engineered proteins.
Our integrated Protein Science and Biophysics team at Concept Life Sciences, are always looking at how they can solve the challenges that science can pose. We understand that timelines are important to all our clients. The successful recruitment of IVTT offers not only a solution for production of toxic proteins, but also for projects where rapid iterative medium throughput cycles are required for protein engineering, such as using AI. Crucially, the developed workflow efficiently delivers active protein for initial SPR analysis.
Pam Dossang, Director of Protein Science, Biophysics and Structural Biology, Concept Life Sciences
The challenge
Typically, recombinant proteins are produced using cell-based expression systems such as bacteria, yeast, insect and mammalian systems. Since these are living systems, production of inherently toxic proteins is not possible. In addition, expression and purification from a cell-based expression system requires longer timescales for each iteration. In general, this offers lower throughput, making it less suited to fast-paced in silico protein design applications.
Our approach
Cell free expression, or in vitro transcription and translation (IVTT), uses cell-derived material to produce proteins in vitro. This enables production of proteins that are typically toxic to living cells in the standard systems used for recombinant protein generation. Furthermore, this approach is highly amenable to investigating multiple constructs in parallel, allowing the production of protein target variants in a multi-well plate format.
Our solution
Protein Expression and Purification
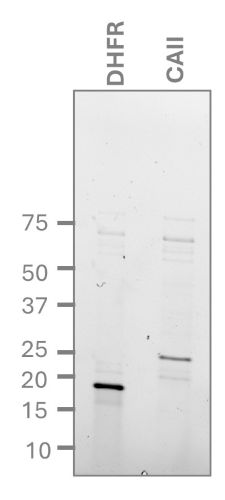
Successful expression was achieved of the human Carbonic Anhydrase II (CAII) with a C-terminal His tag using the NEBExpress® Cell-free E. coli protein synthesis system, under standard conditions. Single step purification was achieved using an NEBExpress® Ni spin column. Protein expression and purification was completed in <5 hours. Fractions were analysed using reducing SDS-PAGE and a final total yield of 18 µg per reaction for CAII protein (Figure 1) was calculated. This yield was sufficient to perform initial SPR analysis for confirmation of protein activity using dihydrofolate reductase (DHFR) as the positive control.
Surface Plasmon Resonance
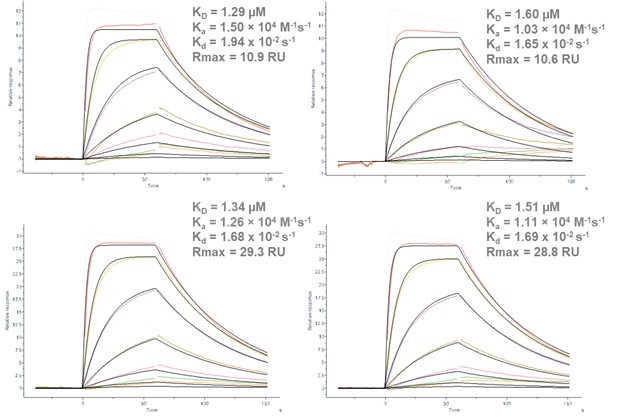
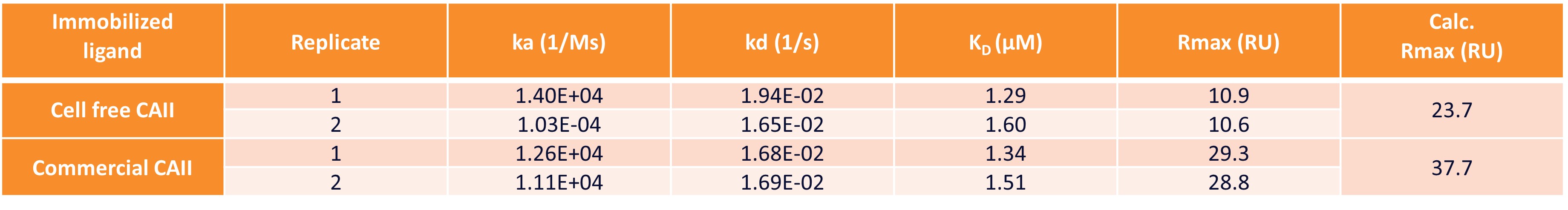
The activity of CAII produced using this cell-free platform was shown to be comparable to commercially available human CAII. The proteins were immobilized on a nitrilotriacetic acid (NTA) surface via amine coupling, which resulted in stable immobilization levels of ~ 2,200 RU (cell-free expressed) and 3,500 RU (commercial positive control). Furosemide, a known small molecule binder of CAII was used as the analyte to assess binding to both proteins using Multi-Cycle Kinetic (MCK) analysis with a half-log concentration series between 31 nM and 100 µM. The generated sensorgrams (Figure 2) were fitted to calculate the affinity and kinetics of this interaction (Table 1).
The result
We delivered active protein of sufficient quality and purity for downstream analysis:
- High purity at microgram level yields.
- Protein activity confirmed by SPR.
- A widely applicable approach for successful initial delivery of toxic proteins.
- A scalable platform for use in medium throughput applications, such as with AI-designed proteins.
